Abstract
Neurons display remarkable sub-cellular specificity in their synaptic targeting, which varies by cell type—for example, excitatory neurons prefer to target the spines of other excitatory cells. Modern electron microscopy connectomes enable the study of this sub-cellular specificity and its context in a circuit at unprecedented scale and resolution. However, this scale has also made it challenging to create accurate and efficient methods for classifying and segmenting fine cell components (including spines) across entire volumes. Here, we present a cost-efficient computational pipeline for classifying postsynaptic targets and segmenting structures such as spines. Our method relies only on having a mesh representation of a neuron and avoids processing image or segmentation data directly. Instead, we leverage tools from geometry processing to create features capturing the local geometry of a neuron’s surface. We couple this core technique with computational and storage optimizations, enabling reliable deployment over hundreds of thousands of neurons for a few hundred dollars in cloud compute cost. We then show that a simple classifier trained on the MICrONS mouse visual cortex dataset can use these mesh-based features to accurately classify synapses as targeting somas, dendritic shafts, or spines (weighted F1 score 0.961). Using this pipeline, we create a map of the postsynaptic structures at over 207.3 million synapses in MICrONS. We present an overview of this census of postsynaptic targeting, finding expected patterns (e.g., excitatory neurons preferentially targeting excitatory spines) as well as unexpected exceptions (e.g., Layer 5 near-projecting and Layer 6 corticothalamic cells often connecting to excitatory neuron shafts). We also demonstrate that these tools can be used to detect spines receiving multiple synaptic inputs, revealing surprising variability in their frequency across cells even within a cell type. We make our postsynaptic target predictions available for study, as well as the code for the computational pipeline and cloud deployment. Beyond MICrONS, we find that the model generalizes well to the H01 connectome without retraining (weighted F1 score 0.949), indicating that these tools will be useful in future connectomics reconstructions. More generally, our work demonstrates that representations derived from neuronal meshes can be a scalable and generalizable primitive for describing morphologies.
Approach: Heat Kernel Signatures on Neuron Meshes
A long-standing challenge in large-scale connectomics is classifying the sub-cellular compartment that each synapse targets---whether it lands on a dendritic spine, a dendritic shaft, or the soma. Prior approaches rely on volumetric image processing or manual annotation, both of which become impractical at the scale of modern millimeter-scale datasets containing hundreds of millions of synapses.
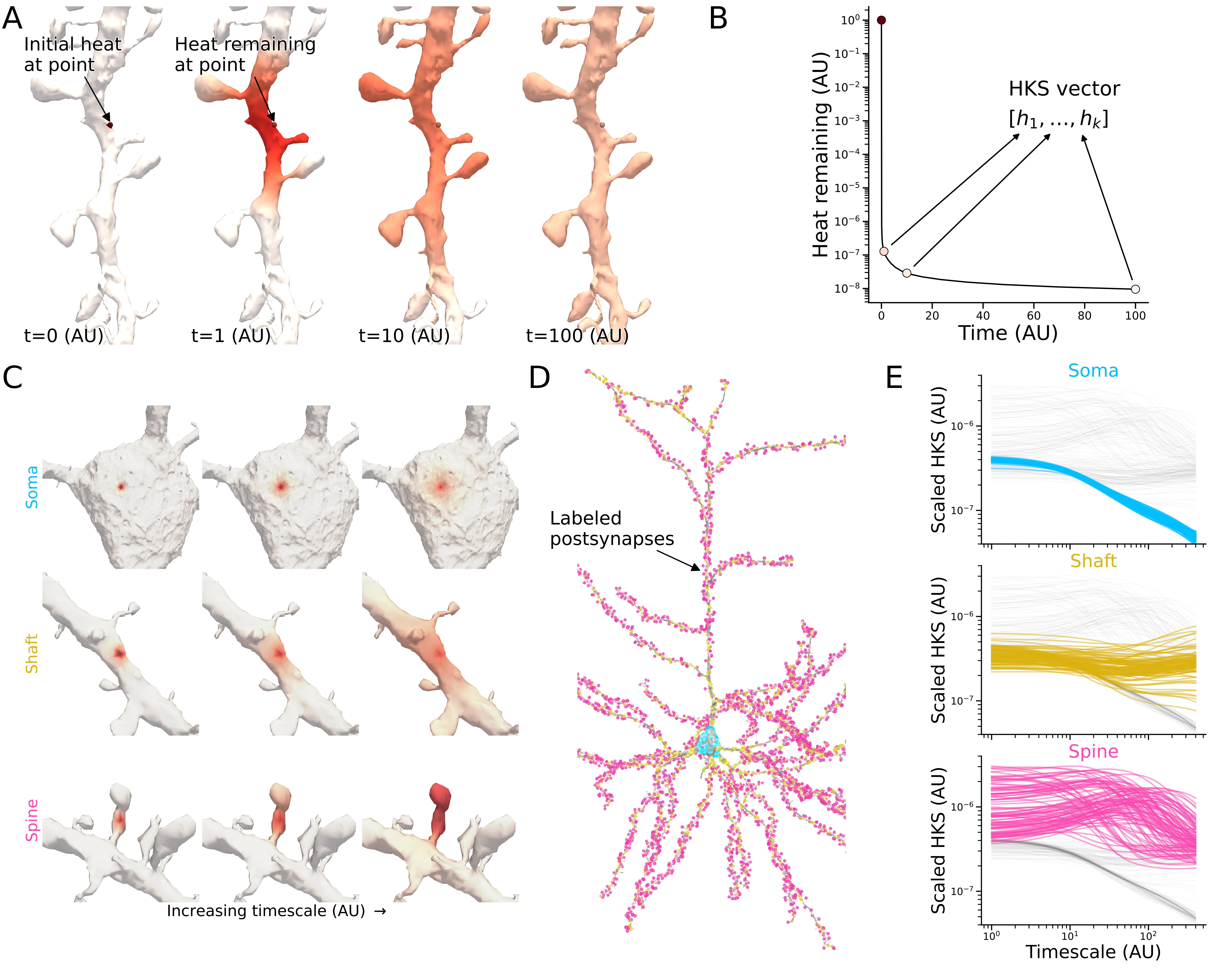
We take a different route: instead of processing the raw imagery, we work directly with the mesh surface of each reconstructed neuron. Specifically, we compute the heat kernel signature (HKS)---a feature derived from how heat diffuses across the mesh surface. Intuitively, a unit of heat placed on an isolated spine is “trapped” and dissipates slowly, whereas heat placed on a broad dendritic shaft or soma spreads quickly across the surface. This differential behavior over a range of timescales produces a compact, discriminative feature vector at each mesh vertex that cleanly separates spines, shafts, and somas without ever touching the underlying electron microscopy imagery.
The HKS is computed from the eigenvectors and eigenvalues of the robust mesh Laplacian:
where and are the -th eigenvalue and eigenvector, and is a timescale. Assembling these values across a range of logarithmically spaced timescales gives a 32-dimensional descriptor at each synaptic site.
A Scalable, Cloud-Deployable Pipeline
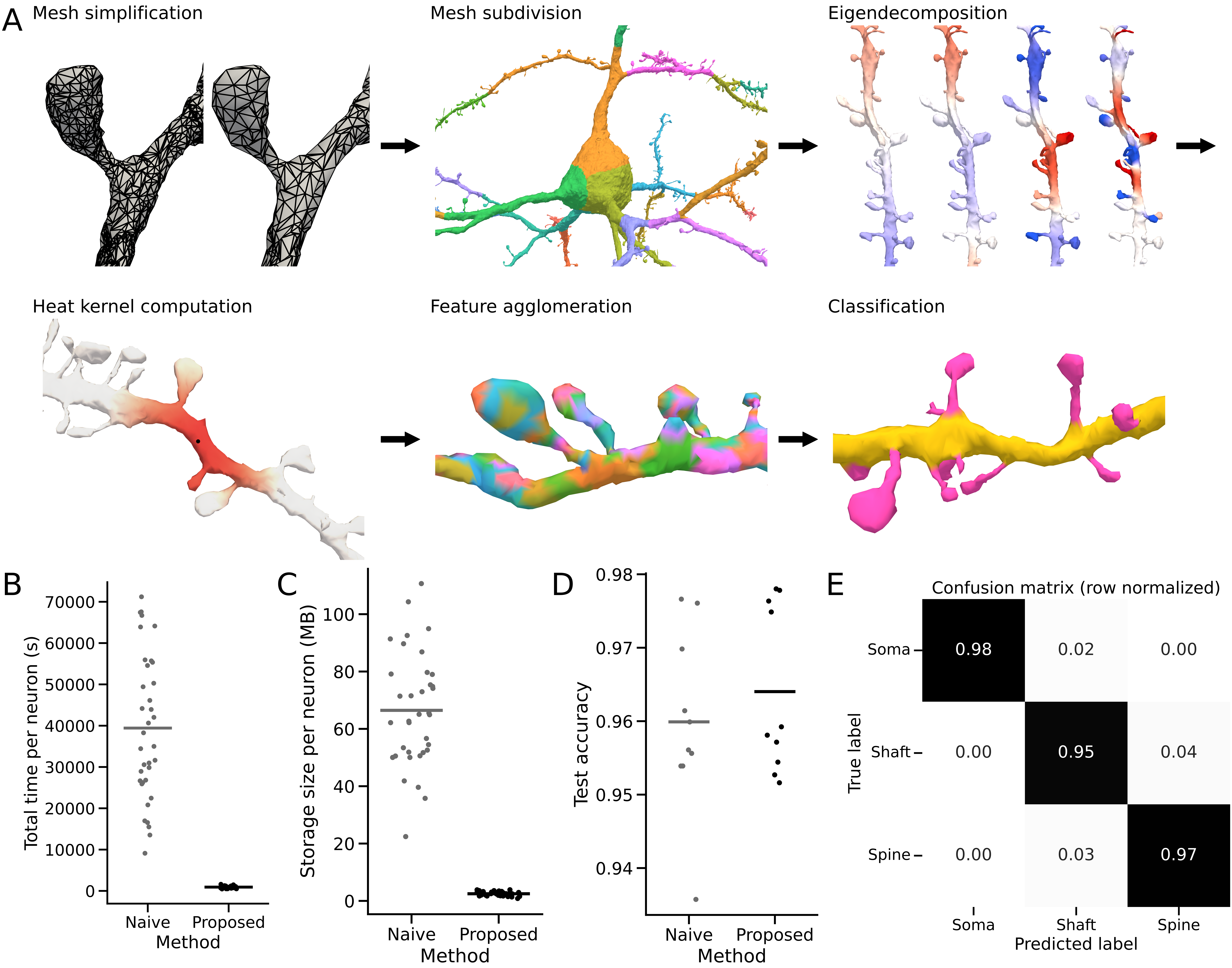
Computing the full eigendecomposition of a neuron mesh with millions of vertices is prohibitively expensive. To scale HKS to an entire cubic-millimeter dataset, we developed a suite of complementary optimizations:
- Mesh simplification via quadric decimation (70% vertex reduction) before eigendecomposition, with negligible accuracy loss
- Overlapping spectral mesh splitting to process large neurons in parallel chunks
- Band-by-band eigendecomposition, computing eigenpairs in batches rather than all at once
- Mesh agglomeration for lossy compression of the HKS vectors, reducing storage by 27× after gzip compression
Together, these yield a 42× speedup in compute time (from ~39,000 s to ~920 s per 36-neuron batch) while maintaining 96.8% classification accuracy---unchanged from the naive implementation. The full pipeline was deployed on the commercial cloud for under $500 to process all 75,241 neurons in the MICrONS dataset.
Interactive Mesh
Classifying 208.6 Million Synapses Across MICrONS
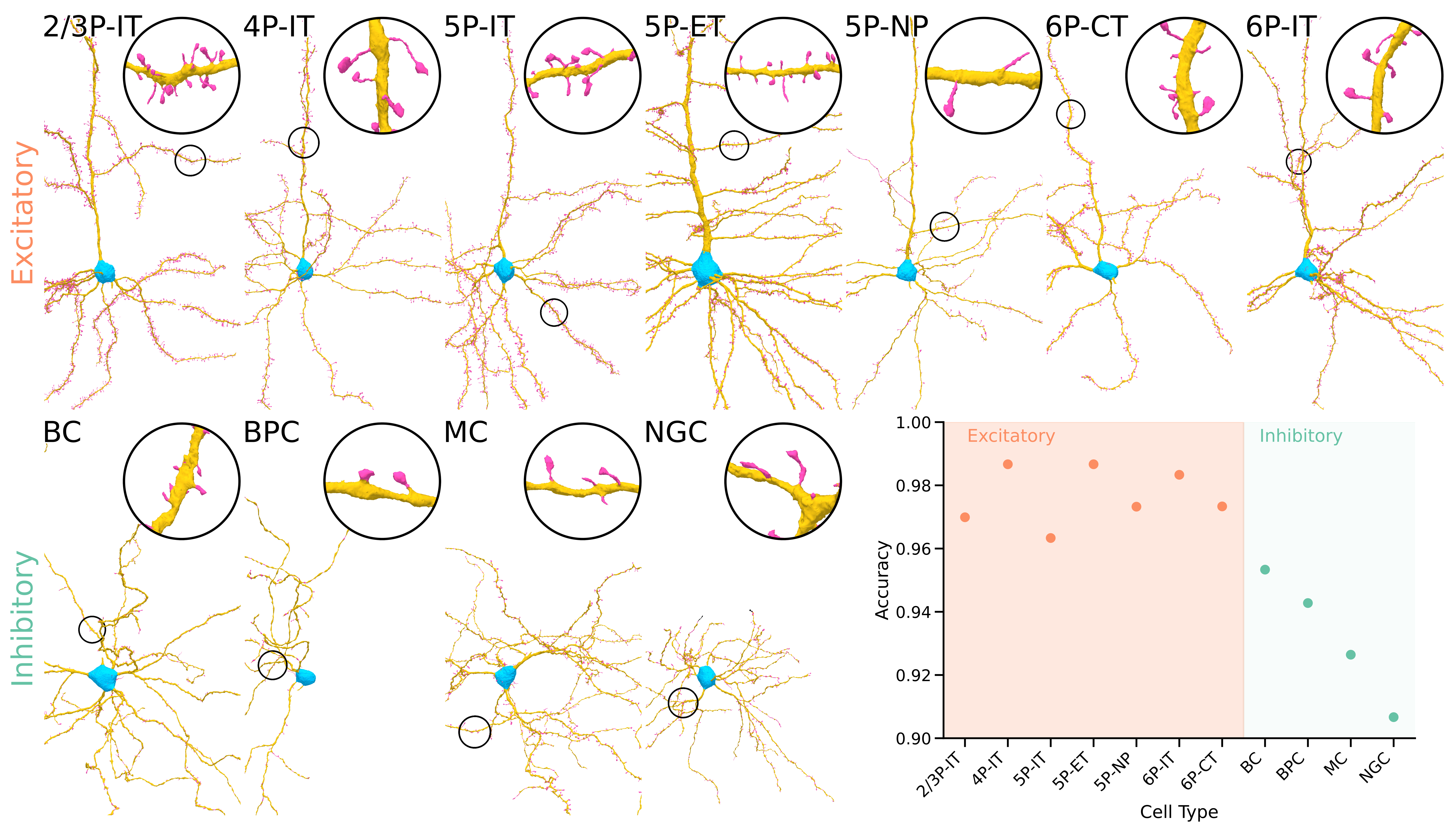
We applied the pipeline to the full MICrONS mouse visual cortex dataset, processing 75,241 neurons and classifying each synapse as targeting a soma, dendritic shaft, or spine. A random forest classifier trained on 110,946 manually labeled synapses achieves a weighted F1 score of 0.961 overall---0.977 on excitatory neurons and 0.925 on inhibitory neurons. This substantially outperforms the NEURD baseline (weighted F1 0.842).
The resulting dataset---postsynaptic target labels for over 208.6 million synapses---is publicly available via the CAVE infrastructure (table: synapse_target_predictions_ssa_v2).
Key Census Findings
Excitatory versus inhibitory targeting. Excitatory neurons in MICrONS receive an average of 70.3% of their synaptic inputs onto spines, compared to just 19.8% for inhibitory neurons. Among excitatory cell types, Layer 2/3 pyramidal cells have the highest spine-input fraction (75.9%) while Layer 5 near-projecting (NP) cells are notably lower (42.1%).
Unexpected output targeting by L5 NP and L6 CT cells. While most excitatory neurons strongly prefer to target spines when contacting other excitatory cells (83.3% on average), Layer 5 NP and Layer 6 corticothalamic (CT) cells are striking exceptions: they direct 45.5% and 44.6% of their excitatory-to-excitatory outputs onto dendritic shafts---a pattern more typical of inhibitory neurons. These two cell types share a late-stage developmental lineage, suggesting a common developmental program may underlie this unusual targeting preference.
Multi-input spines are highly variable. The pipeline also enables detection of spines that receive more than one synaptic input. On average, 6.1% of excitatory neuron spines are multiply innervated---but this fraction is strikingly variable: among Layer 2/3 pyramidal cells alone, the range spans from 2.2% to 19.3% across individual cells. This variability cannot be explained by spine size, soma depth, total synapse count, or other measured morphological features, pointing to dynamic processes not captured by the static connectome.
Inhibitory inputs preferentially target multi-input spines. Thalamic and local excitatory axons direct 10.8% and 8.2% of their outputs to multi-input spines, respectively. Inhibitory neurons, by contrast, send approximately 70% of their outputs to multi-input spines---far exceeding their representation. Multi-input spines are also significantly larger than single-input spines, with a strong correlation between spine volume and excitatory cleft size () but only a weak correlation with inhibitory cleft size ().
Data & Code
BibTeX
@article {pedigo_quantitative_2026, author = {Pedigo, Benjamin D. and Danskin, Bethanny P. and Swanstrom, Rachael and Neace, Erika and Dorkenwald, Sven and da Costa, Nuno Ma{\c c}arico and Schneider-Mizell, Casey M. and Collman, Forrest}, title = {A quantitative census of millions of postsynaptic structures in a large electron microscopy volume of mouse visual cortex}, elocation-id = {2026.02.19.706834}, year = {2026}, doi = {10.64898/2026.02.19.706834}, publisher = {Cold Spring Harbor Laboratory}, URL = {https://www.biorxiv.org/content/early/2026/02/20/2026.02.19.706834}, eprint = {https://www.biorxiv.org/content/early/2026/02/20/2026.02.19.706834.full.pdf}, journal = {bioRxiv}}